 |
DNA Sequence Assembler
Release history
What's new in version DNA Sequence Assembler 5?
- New: 64 bit support. DNA Sequence Assembler can now assemble much much longer contigs! The amount of available RAM is now the only limitation.
- New: Added 'Highlight IUPAC bases' function in grid
- New: Added 'Show original bases' function in chromatogram
- New: Added 'Show original bases' function in grid
- New: View RC of contig in dedicated viewer
- New: ABI: More fields/info presented in 'ABI properties'.
- New: ABI: Tags presented in 'ABI properties'.
- New: Allow user to save all chromatograms to disk.
- New: File name shown at the beginning of the chromatogram ..
- New: MultiFASTA files are open in an external/dedicated viewer.
- New: MultiGbk files are open in a dedicated viewer.
- Improved: MUCH better handling for corrupted SCF/ABI files
- Improved: Improved QV assesment for SCF files
- Improved: Alignment window: control auto-alignment now also works for projects with lots of sequences (control alignment)
- Improved: Assembly log is now shown next to the assembly grid
- Improved: Assembly window caption
- Improved: Batch assembly by subfolders. now the name of the contig is composed from the name of the current sub-folder
- Improved: Batch assembly by subfolders: don't show the 'ADD/REMOVE files' buttons in Project Builder
- Improved: Function 'Center cursor on screen' renamed to 'Extended selection'
- Improved: Change IUPAC base background color to make text more readable
- Improved: Function 'Copy contig to clipboard': The recognition sequence is automatically removed from contig
- Improved: Function 'Highlight contig errors' renamed to 'Highlight mismatches'
- Improved: Function 'Highlight IUPAC bases in chromatogram'
- Improved: Function 'Highlight low quality bases'
- Improved: Function 'Highlight mismatches'
- Improved: Function 'Use rainbow colors' was improved
- Improved: hint in the pop-up window that appears when the cursor is placed over a base
- Improved: mismatches count
- Improved: The size of Assembly window is always a bit too big/small
- Improved: The words 'Mutation mode' appears not in Chromatogram Editor when it is running in 'Mutation mode'
- Improved: Toolbar in Assembly window
- Improved: Faster start up
- Improved: Faster program shutdown
- Improved: ABI: Abi importer massively improved (better handle malformed files).
- Improved: ABI: lower memory footprint when loading ABI files..
- Improved: Better tool-tips.
- Improved: Fixed some English typos.
- Improved: Global importer improved (better handling of malformed files).
- Improved: GUI improvements.
- Improved: Improved chromatogram management.
- Improved: Improved chromatogram renderer.
- Improved: Improved message: reason why a sequence could not be assembled.
- Improved: Improvements to FASTA/multiFASTA parser.
- Improved: Improvements to FASTA/multiFASTA viewer.
- Improved: Improvements to 'Sequence file dereplicator'.
- Improved: Internal Sequence Viewer now also shows GC percent.
- Improved: Multiple improvements to Log.
- Improved: Reference marked in special color when the contig contains a mismatch/mutation. (Benjamin).
- Improved: 'Reverse complement file' was added (additionally to the already existent 'Reverse complement sequence').
- Improved: SCF files: The original comment is kept when a copy of the SCF is created.
- Improved: The 'Batch sequence assembly by filename pattern' module was dramatically improved.
- Improved: The 'Tasks' were renamed to 'Projects'.
- Fixed: An error is shown while loading a SCF file that was save while in mutation mode. This is because the ambiguous bases were converted from ACGT to IUPAC.
- Fixed: Correctly display chromatograms that are too short.
- Fixed: Fixed: MRU (most recent used) menu!
- Fixed: In some cases the last few samples of a chromatogram were not copied to clipboard.
- ... many other fixes and improvments.
- Note: Projects saved with DNA Sequence Assembler v4 can still be loaded in DNA Sequence Assembler v5.
What's new in version DNA Sequence Assembler 4?
- Mutation detection module added
- The following sub-modules were added:
- Single organism (Multiploid, Diploid) mutation detection
- Different organisms (Haploids, Multiploids, Diploids) mutation detection
- The user can jump to Next Mutation and to Previous Mutation with Shift+Click
- Added 'Use internal base caller to calculate quality values' checkbox (allows the user to enable/disable the internal base caller)
- Added 'Detect mutations' checkbox on interface
- Added 'Mutation detection parameters' on interface
- The user can choose to show N bases as mutations.
- Log: Show parameters used for mutations in log
- Log: Single viewer - Show total mutations, position and base for each mutation
- Log: Number after mutation not after sequence
- Log: Mutation different organisms - Show total mutations, position and base for each mutation
- Log: Mutation same organism - The report is the same as for Mutation different organisms but look first at the rules for the contig assembly
- Log: In Mutation mode the Log will show the ratio between areas of two overlapping bases
- Log: Show base numbering based on contig/reference/ruler
- Log: In v4.9 the log was dramatically improved. Now mutations are shown in a table. This helps the user to quickly overview the results
- New 'Highlight mutations' button added. It has effect on all open chromatograms and contigs.
- Mutations are highlighted in purple color in contig
- IUPAC bases entered by the user (in contig) are highlighted in pink
- State of the art base caller
- Blazing-fast proprietary algorithm that uses a sum of 7 different algorithms to compute the data accurately. Used to determine secondary peaks, uncalled peaks, and QV info.
- Single nucleotide polymorphism (SNP) detection module added
- The new advanced SNP detection offers one of the most accurate solutions on the market
- Secondary peak detection added
- SNPs are shown now as IUPAC code
- Use ambiguity code for mismatches only when the QVs are similar
- Major improvements in the sequence assembler
- DNA Sequence Assembler now makes more accurate suggestions for contig ambiguities. It chooses the base with highest QV. If the input samples do not have QV info embedded OR if all bases have the same QV, then it chooses the base that appears most often.
- Automatic detection of sequence/contig direction (forward/reverse) based on its filename pattern
- New consensus algorithm allows the user to assemble QV and non-QV files together (for example a FASTA files and a SCF file) while still generating accurate contigs
- Automatic homopolymers error correction added.
- New option (checkbox) added: 'FASTA files are trusted'. In Mutation mode the Fasta files are always trusted because it is considered that the checked file was used before starting the mutation detection process
- For flexibility, 'Minimum overlap' can be set to a much lower value
- The GAP insertion algorithm was improved
- Major improvements in the automatic end trimming module
- The automatic end trimming (cleaning low quality regions that the end of the sample) was extended and improved for accuracy
- Automatically cut bad ends (NNNNNNNNNNNNNNs) in files that contain no quality scores (such FASTA).
- Major improvements in the Chromatogram Editor
- Show link between bases and chromatogram peaks
- The user can now click a base to select/highlight it
- New button added to show/hide A, C, G, T traces independently
- Copy entire chromatogram to clipboard
- The user can choose if the chromatogram will show 'base letter', 'QV', or 'base position' next to each peak
- Better scroll bar
- The Chromatogram Viewer can handle now invalid (too short (less than 10 bases)) chromatograms
- Improved 'Contig Assembly' window
- Better base selection. The user can start a selection (click and drag) even if the starting base was already selected.
- Better hints
- Better menus
- 'Use rainbow color', 'Keep cursor on center', 'Highlight low quality bases' options are now accessible also from the main menu
- When a base is clicked in Chromatogram Viewer the Assembly grid jumps also to that base (sync editing)
- Focus line (dotted line) inside the focused cell is not shown anymore.
- The program now remembers user's settings for Assembly window
- Show confirmation that the contig was saved to disk when the user pressed the 'Save contig' menu/button
- Allow user to 'Optimize assembly for very low quality samples'
- New buttons for Next/Previous Error
- The 'Base editor' is shown when the user presses Enter key (initially worked only with double click)
- Better Project Manager
- Added 'Mutations' tab in Project Manager. The user can change here the parameters for mutation detection
- Added 'Browse for reference file' button in Project Builder tab
- The Project Manager can now also show 'Hidden' folders
- Better 'project name' auto-generation. The user can stop the program for auto-generating names.
- In 'Sample Explorer', the 'All supported files' filter now also includes 'BaserProjects' files
- The 'DNA Sequence Assembler Projects' filter was moved two lines up, after 'All Supported Files'
- 'Sequence processing options' tab -> The 'Auto open this tab on assembly failure' option appears only for the assembly jobs
- Accessibility improvements
- The keyboard shortcuts have been fully redesigned
- Added a button in the main toolbar for easy access to Project Manager.
- When the contig assembly is done the cursor is automatically moved to the first ambiguous base in contig
- The job type (batch, single contig, viewer, mutation detection, etc) is shown in Assembly Window's caption so the user can easily see it
- The Vectors window moved in the Project Manager for faster access
- Added link to 'TheScientist.info' in the Info menu
- Added 'Load sample' button in Sequence Viewer
- The 'You can double click a sample to view it' message is shown when the user clicks 'View single chromatogram'.
- Added menu for easy access to SFF Workbench external tool
- Added 'Save contig as RC (reverse complement)' menu under ‘Contigs‘
- Added "How to cite DNA Sequence Assembler " menu
- The 'Demo samples' folder added to the 'MRU' (most recent used) so the beginners easily access it when DNA Sequence Assembler starts for the first time
- Added ‘Tasks’ button also in Project Manager for easier access.
- Close Tasks on DoubleClick on its caption
- Tutorial messages that suggest to the user to use the F9 keyboard shortcut for ‘Start job’
- When the user presses the ‘Save contig’ button let it know that the contig was saved
- New shortcut added. By pressing F9 key the user can perform the current selected action (assemble, batch assemble, convert to Fasta, etc)
- Let user always choose if the program starts in 'Assembly mode' or 'Last used mode'
- Improved file handling
- Better reports (log) when converting SFF to FASTA
- The 'Batch file convertor' tool can now use the filename as comment
- Export all samples at once as SCF.
- DNA Sequence Assembler v4 can still open DNA Baser Assembler v3 projects
- Improved registration process
- New, less intrusive registration system
- DNA Sequence Assembler was split into modules. The user can purchase all modules or a limited set of modules (at a smaller price)
- The user will get a new trial period (to reevaluate the program) when a new version is released
- New 'Price calculator' module integrated directly into the program
- New uninstaller is available
- The Price Calculator was embedded in DNA Sequence Assembler now. The user can now purchase DNA Sequence Assembler directly from the program.
- Major improvements in the graphic interface
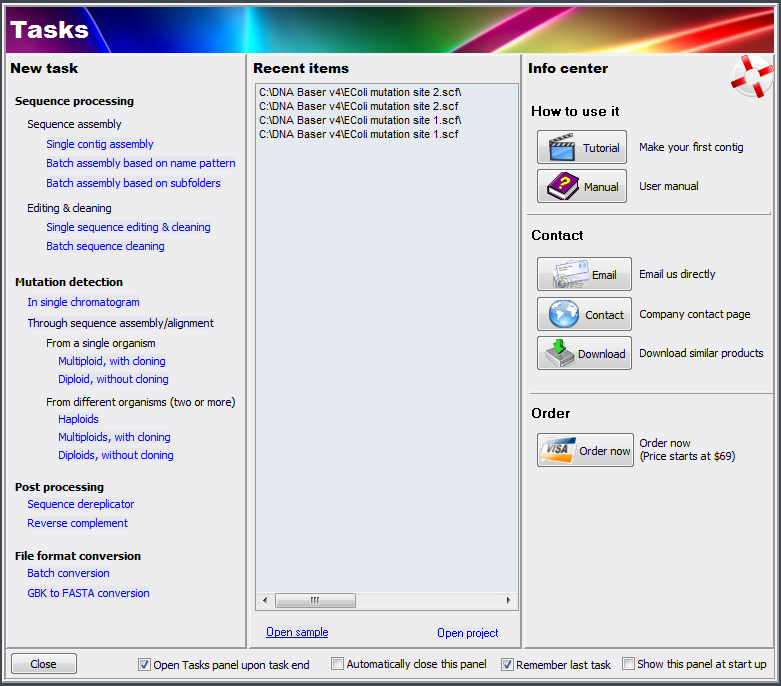
- The GUI workflow was also massively improved. The interface in now based on 'Tasks' (see image below)
- New 'Chromatogram Preview'
- Allows you to quickly asses the chromatograms as you work with them. It also shows statistics for the current selected file.
- It can also show the CG and AT percent
- Mini-chromatogram overview
- New checkbox in Task Manager: 'Open Tasks panel when current task ends'. If enabled, every time the user closes a tool (batch processing, Assembly window, form manager, etc) the Tasks Manager will appear.
- New pop-up menu on the main 'Start action' button allows quick access to predefined assembly settings
- New 'Info Center'
- Better menu icons
- Show MRU (Most Recent Used) files in the Tasks panel
- Improved 'Search again' function: if the user presses 'Search again' while the search box is empty, the 'Search' panel will show up.
- Improved 'Save as...' dialogs (v4.5)
- New 'Browse' button added in Sample Explorer (v4.2)
- Tutorial for 'Sequence editing' module added (v4.5)
- Improved 'Browse folder' dialog
- Better info about the progress of the assembly job(s)
Other improvements
- 'Remember last edited' base was improved
- When the assembly process is done, scroll to the first ambiguous base
- The Mutation Log was moved in its own tab. Show/Hide tab as necessary. 'Mutations' tab is shown only in mode 2 and 5 (v4.10)
- Auto-generate project name & path' in now check by default (v4.5)
- Improved Chromatogram Viewer - The empty area at the beginning/end of a chromatogram is indicated with a diagonal pattern (v4.5)
- Better log in Sequence Dereplicator (v4.5)
- The 'Recompute N bases' feature was separated from 'Detect Double Peaks' feature
- When the Project Manager is in 'Batch seq cleaning' mode, the 'Name' edit box is made invisible (it is not needed)
- The Assembly window now rememebers the last status of 'Info Line' combo box
- The user is not allowed anymore to click in an empty area in a Chromatogram
- Locate sample' menu added in Assembly window. When clicked, DNA Sequence Assembler will open Windows Explorer to the folder that contains the sample
- 'View sample alone' menu added in Assembly window. When clicked, DNA Sequence Assembler will load that sample alone in SampleViewer
- The user can move the Tasks window
- Added a cehckbox next to 'Project path'. Used to instruct the program to keep existing path or to generate new paths as the user navigates through folders
- Improved (Win 7 style) dialog boxes
DNA Sequence Assembler v3
- Save the entire project to disk (samples, contig, user editing and corrections, etc). It can be done manually and/or automatically.
- Edit existing projects (Add/delete samples to project)
- Remember the edits that the user made in contig/input samples when the contig is recalculated.
- MRU (Most Recent Used Files and Folders) - Quickly re-open a sample file/project that was used recently.
- Batch by name pattern based on separator (thanks to Stephan)
- Tutorial panel (with animated logo and sound on new announcement). When the program starts for the first time it starts in tutorial mode
- New auto-updater tool
- Automatically start DNA Sequence Assembler when a project file is double clicked in Windows Explorer
- Added separate popup menu when the user clicks on Assembly Grid header (it contains 'Delete current sample', 'Sample properties', 'Save sample as', etc)
- Added 'Sample' menu in Contig Assembly window
- Added 'Project' menu in Contig Assembly window
- Added 'Delete column' function
- Added 'Create new FASTA sample' tool
- Added 'Save contig as RC' menu
- Added 'Save contig' button in Assembly window
- Added "View log" button in Assembly window.
- Added several new menus in ‘View’ main menu
- Added icons for all tabs in Project Manager (16x16 and also 32x32 pixels)
- Added top logo in Project manager
- Added 'Mark as trusted/untrusted' button in Assembly window
- Added 'Ruler offset' feature (under 'Project' menu)
- The user can start the assembly even if there is a single multiGbk/multiFasta file in Project Manager
- The caption of Assembly window shows the selection size (how many bases were selected by user)
- The user can now stop the single contig assembly and batch contig assembly process by pressing the Esc key
- Let the user drag and drop files directly into DNA Sequence Assembler main window. This will not add the files to the job list but it will make the Project Manager to navigate to the folder that contains the dragged files.
- Added 'Export alignment as text file' function which saves the entire alignment to disk as text file
- The user can scroll the Assembly grid from Contig Map (by dragging the blue selection)
- The user can change the active sample in the Assembly grid from Contig Map (by clicking a sample)
- 'Rainbow' colors for assembly grid background - Helps you to quickly stop ambiguities, mismatches, errors
- New: Show 'Search not found' message when the searched sequence was not found.
- New: Put a bunch of imported sequences in a list (feature requested by lcrerar/masonlive.gmu.edu)
- New: The program is now monolithic (you can copy only the EXE file on a USB stick and take it with you).
- Clicking on sample's name (first column in Assembly grid) makes the cursor to jump at the beginning of that sample
- Hovering the mouse over Assembly grid's first column will show not only the file name of the sample but also its comment
- For multi-FASTA files the first column of the Assembly grid shows the name of that file followed by incrementing numbers (one for each multi-FASTA sub-component)
- 'Sequence recognition' window was improved
- Move the "Automatically switch to log" checkbox under the log.
- Some menu re-arrangement
- Speed improvement
- Many other graphic user interface improvements (such as colors in grid and contig map, layout rearrangement, etc)
- Added (partial) support for samples that contain lowercase bases.
- The level of trust for current contig is now reported in Log
- Totally rewritten selection tool. The selection follows the cursor as it jumps between rows.
- Improved: Better Network package.
- Improved: Dramatic improvements related to memory requirements have been made. Now you can assemble very long samples (at least two times longer than before) without running out of memory.
- Improved: Don't let the user put the selection in the empty (no bases) area.
- Improved: DNA Sequence Assembler can open very large multiFasta files now
- Improved: Batch assembly FASTA/SEQ/GBK sample viewer
- Improved: Drag and drop file directly into the main form will open the file
- Improved: Fasta2MultiFasta converter updated to open all supported file types (scf, abi, gbk, etc)
- Improved: MultiFasta2Fasta converter updated to open all supported file types (scf, abi, gbk, etc)
- Improved: Sequence assembly speed was dramatically improved (up to 30% in some cases)
- Improved: Added 'Do nothing' option in 'File operation for invalid/unassembled files' in Project Manager
- Improved 'Batch by subfolders' and 'Batch by filename pattern'
- Improved: Batch mode: The program is now checking if it can create output folders
- Improved: Better Unicode support.
- Improved: Loading text samples (Fasta, Seq) requires less RAM now. No (pseudo) chromatogram is created for simple text samples (Fasta, Gbk, Txt, Seq).
- Improved: Log (Make log compatible with DNA Sequence Assembler Console. Save all text in log to disk (save project) and restore it later)
- Improved: Message: "Information about contig is shown only if the 'Auto save contig to disk' (located in 'Project Options' tab) is active."
- Improved: If there are unassembled multi-Fasta files show this message: "Samples belonging to multi-Fasta/multiGBK files will not be moved to Unassembled folder"
- Improved: The vertical scroll-bar in Contig Map appears now only when necessary.
- Improved: GUI is now supporting Windows 7 themes:
- GUI: Contig bases that will not be saved to disk (because vector removal) are stroked out
- GUI: The layout of the Assembly window was improved. One of the improvements is the search box which is now bigger and detached
- GUI: Less flicker during program start up
- GUI: More descriptive text for main buttons
- GUI: The tool tips pop-up faster now
- GUI: Better log for Batch Sample Processing
- GUI: Ambiguities are shown in a different shade of red (different than low confidence score bases)
- GUI: For better visibility the current selected sample is now indicated with an arrow and its background is yellow.
- GUI: The reference sample is shown in pink color also in Contig Map
- Better support for ABI files
- Faster Job List (it was slow when lots of samples were loaded into it)
- Untrusted ends are shown in dark shades also in reference
- The log now show the total number of invalid samples
- The Home/End keys moves the cursor to the beginning/end of current sample instead of beginning/end of Assembly Grid
- The window size and screen position for 'FASTA Viewer' is remembered next time when the program starts
- In the Assembly window the tool-tip now shows two values for the position of the base: the position counting the gaps introduced by the assembly process and the position without these gaps
- Automatically associate DNA Sequence Assembler with project files.
DNA Sequence Assembler v2
- Let user resize chromatograms
- Support for SEQ, FASTA, TXT files
- Support for SCF v3.10
- Contig map
- Drag and drop support from Windows Explorer
- Accept command line parameters
- Batch sequence assembly
- Other features...
DNA Sequence Assembler v1.2 (Nov. 2005)
- Added support for ABI, SCF
- Sequence recognition (primer/vector) trimming
- Let user insert a base
- Let user delete bases from contig
- Show edits in green color
- When moving cursor over a base, show base info
- Other features...
 Building software together Building software together
DNA Sequence Assembler is more user-friendly and more affordable than any other similar product on the market. It could not be this way without help from our customers and beta testers. By submitting feature requests, feedback and bug reports you help us to make a better product and to keep its price as low as possible. At the end you will directly benefit from it.
Please don't hesitate email us or to chat (live) with us! Your opinion matters!
Stay tuned
Do you want to receive news as new updates are released? Just send us an email with the subject "subscribe".

|
 |
